Principled Bayesian Workflow
On Thursday evening Michael Betancourt gave an insightful and thought provoking talk on Principled Bayesian Workflow at the Baysian Mixer Meetup, hosted by QuantumBlack.
Michael is an applied statistician, conslutant, co-developer of Stan and passionate educator of Bayesian modelling.
What is a principled Bayesian workflow?
It turns out that it mimics my idea of the scientific method:
- Create a model for the ‘small world’ of interest, i.e. the small world relevant to test an idea, e.g. law of gravity that describes how an apple falling off a tree is accelerated by the Earth’s gravitation
- Decide which measurements will be relevant
- Make predictions with the model
- Review if the predictions are reasonable
- Set up experiment and predict outcome with set parameters
- Carry out experiment
- Compare measurements with predictions and go back to 1 if necessary
Michael summarised his approach more eloquently for Bayesian Inference in one slide: 
He gave also succinct advise for model calibration and validation. Michael suggested the following four questions:
- Are modelling assumptions consistent with our domain knowledge?
- Are our computational tools sufficient to accurately fit the model?
- How should we expect our inference to perform?
- Is our model rich enough to capture the relevant structure of the data generating process?
Adding structure could mean to add more parameters to the model, rather than changing it fundamentally. As an example Michael mentioned a model that started with a Normal distribution for the data generating process, but that was expanded to a Student’s t-distribution to allow for fatter tails. The reasoning here is that Student’s t-distribution will converge to \(\mathcal{N}(0,1)\) for \(\nu \to \infty\) and hence, incorporates the initial assumption of normality.
From his experience it is helpful to show data (including data generated from your model), when asking for advise from domain experts. Experts are more likely to have a view/ opinion on data rather than parameters, and even more so what data will be unreasonable. Don’t forget they are experts, and hence like/likely to disagree with the non-expert. Use this bias to your advantage.
Finally, he showed one chart to check the posterior predictive distribution that I found very insightful. You plot the posterior z-scores against the posterior shrinkages (\(s\)), which are defined as:
\[ \begin{aligned} z & = \left| \frac{\mu_{\mbox{post}} - \tilde{\theta}}{ \sigma_{\mbox{post}} } \right| \\ s & = 1 - \frac{\sigma^{2}_{\mbox{post}} }{ \sigma^{2}_{\mbox{prior}} } \end{aligned} \]
Depending on where you see a cluster in the diagram, it will provide guidance on where you might have a problem with your model. 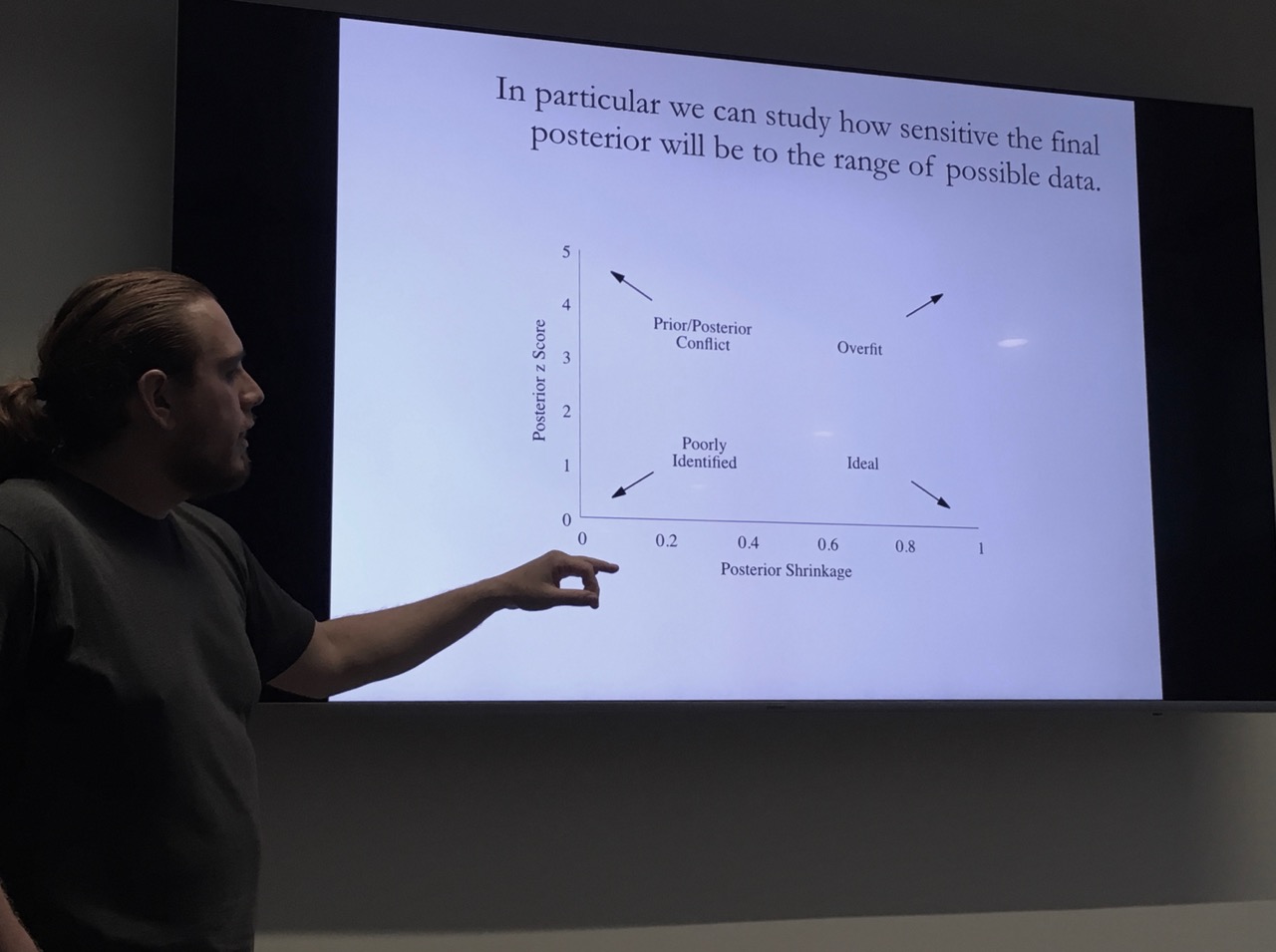
You find Michael’s full article on principled Bayesian workflow on his website.
Next events
The next Bayesian Mixer will take place on 17 July at SkillsMatter. Eric Novik (CEO, Generable) will talk about Precision Medicine With Mechanistic, Bayesian Models.
In addition, on 16 July we have the Insurance Data Science Conference at Cass Business School London, followed by a Stan Workshop on 17 July. You can register for either event here.
Citation
For attribution, please cite this work as:Markus Gesmann (Jun 18, 2018) Principled Bayesian Workflow. Retrieved from https://magesblog.com/post/principled-bayesian-workflow/
@misc{ 2018-principled-bayesian-workflow,
author = { Markus Gesmann },
title = { Principled Bayesian Workflow },
url = { https://magesblog.com/post/principled-bayesian-workflow/ },
year = { 2018 }
updated = { Jun 18, 2018 }
}