Transforming subsets of data in R with by, ddply and data.table
Transforming data sets with R is usually the starting point of my data analysis work. Here is a scenario which comes up from time to time: transform subsets of a data frame, based on context given in one or a combination of columns.
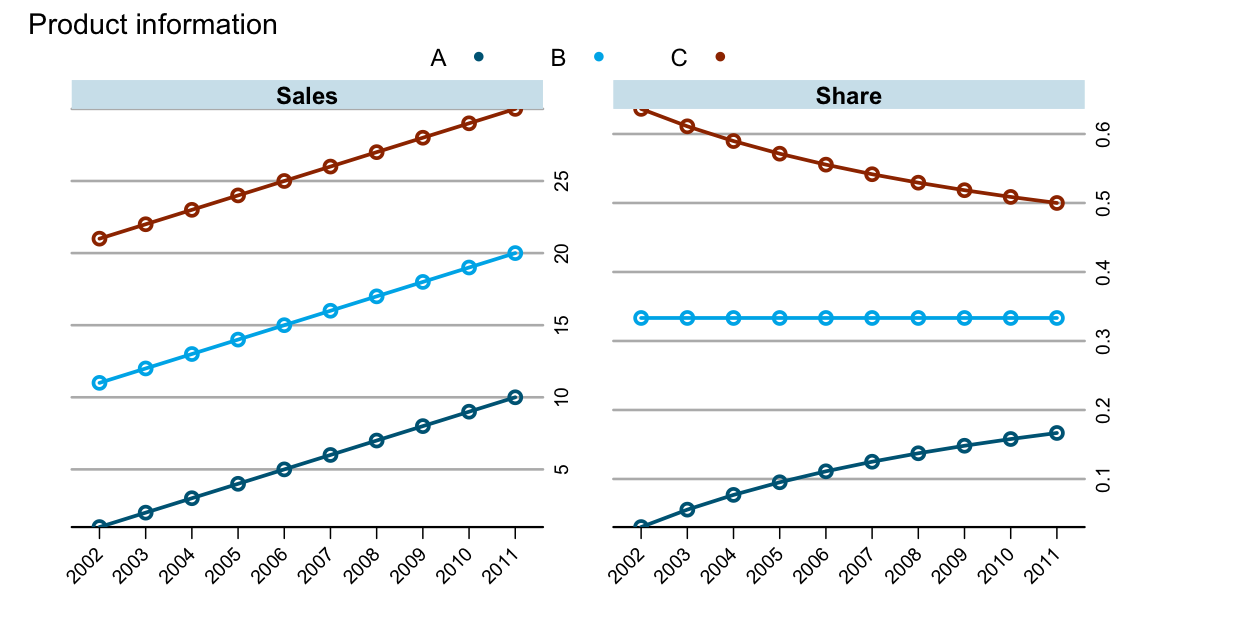
As an example I use a data set which shows sales figures by product for a number of years:
df <- data.frame(Product=gl(3,10,labels=c("A","B", "C")),
Year=factor(rep(2002:2011,3)),
Sales=1:30)
head(df)
## Product Year Sales
## 1 A 2002 1
## 2 A 2003 2
## 3 A 2004 3
## 4 A 2005 4
## 5 A 2006 5
## 6 A 2007 6
- base R with
by, ddplyof theplyrpackage,data.tableof the package with the same name.
by
The idea here is to use by to split the data for each year and to apply the transform function to each subset to calculate the share of sales for each product with the following function: fn <- function(x) x/sum(x). Having defined the function fn I can apply it in a by statement, and as its output will be a list, I wrap it into a do.call command to row-bind (rbind) the list elements:
R1 <- do.call("rbind", as.list(
by(df, df["Year"], transform, Share=fn(Sales))
))
head(R1)
## Product Year Sales Share
## 2002.1 A 2002 1 0.03030303
## 2002.11 B 2002 11 0.33333333
## 2002.21 C 2002 21 0.63636364
## 2003.2 A 2003 2 0.05555556
## 2003.12 B 2003 12 0.33333333
## 2003.22 C 2003 22 0.61111111
ddply
Hadely’s plyr package provides an elegant wrapper for this job with theddply function. Again I use the transform function with my self defined fn function:library(plyr)
R2 <- ddply(df, "Year", transform, Share=fn(Sales))
head(R2)
## Product Year Sales Share
## 1 A 2002 1 0.03030303
## 2 B 2002 11 0.33333333
## 3 C 2002 21 0.63636364
## 4 A 2003 2 0.05555556
## 5 B 2003 12 0.33333333
## 6 C 2003 22 0.61111111
data.table
With data.table I have to do a little bit more legwork, in particular I have to think about the indices I need to use. Yet, it is still straight forward:library(data.table)
## Convert df into a data.table
dt <- data.table(df)
## Set Year as a key
setkey(dt, "Year")
## Calculate the sum of sales per year(=key(dt))
X <- dt[, list(SUM=sum(Sales)), by=key(dt)]
## Join X and dt, both have the same key and
## add the share of sales as an additional column
R3 <- dt[X, list(Sales, Product, Share=Sales/SUM)]
head(R3)
## Year Sales Product Share
## [1,] 2002 1 A 0.03030303
## [2,] 2002 11 B 0.33333333
## [3,] 2002 21 C 0.63636364
## [4,] 2003 2 A 0.05555556
## [5,] 2003 12 B 0.33333333
## [6,] 2003 22 C 0.61111111
data.table may look cumbersome compared to ddply and by, I will show below that it is actually a lot faster than the two other approaches.Plotting the results
With any of the three outputs I can create the chart from above with latticeExtra:
library(latticeExtra)
asTheEconomist(
xyplot(Sales + Share ~ Year, groups=Product,
data=R3, t="b",
scales=list(relation="free",x=list(rot=45)),
auto.key=list(space="top", column=3),
main="Product information")
)
Comparing performance of by, ddply and data.table
Let me move on to a more real life example with 100 companies, each with 20 products and a 10 year history:
set.seed(1)
df <- data.frame(Company=rep(paste("Company", 1:100),200),
Product=gl(20,100,labels=LETTERS[1:20]),
Year=sort(rep(2002:2011,2000)),
Sales=rnorm(20000, 100,10))
I use the same three approaches to calculate the share of sales by product for each year and company, but this time I will measure the execution time on my old iBook G4, running R-2.15.0:
r1 <- system.time(
R1 <- do.call("rbind", as.list(
by(df, df[c("Year", "Company")],
transform, Share=fn(Sales))
))
)
r2 <- system.time(
R2 <- ddply(df, c("Company", "Year"),
transform, Share=fn(Sales))
)
r3 <- system.time({
dt <- data.table(df)
setkey(dt, "Year", "Company")
X <- dt[, list(SUM=sum(Sales)), by=key(dt)]
R3 <- dt[X, list(Company, Sales, Product, Share=Sales/SUM)]
})And here are the results:
r1 # by
## user system elapsed
## 13.690 4.178 42.118
r2 # ddply
## user system elapsed
## 18.215 6.873 53.061
r3 # data.table
## user system elapsed
## 0.171 0.036 0.442
It is quite astonishing to see the speed of data.table in comparison to by and ddply, but maybe it shouldn’t be surprise that the elegance of ddply comes with a price as well.
Addition (13 June 2012): See also Matt’s comments below. I completely missed ave from base R, which is rather simple and quick as well. Additionally his link to a stackoverflow discussion provides further examples and benchmarks.
Finally my session info:
> sessionInfo() # iBook G4 800 MHZ, 640 MB RAM
R version 2.15.0 Patched (2012-06-03 r59505)
Platform: powerpc-apple-darwin8.11.0 (32-bit)
locale:
[1] C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] latticeExtra_0.6-19 lattice_0.20-6 RColorBrewer_1.0-5
[4] data.table_1.8.0 plyr_1.7.1
loaded via a namespace (and not attached):
[1] grid_2.15.0
Citation
For attribution, please cite this work as:Markus Gesmann (Jun 12, 2012) Transforming subsets of data in R with by, ddply and data.table. Retrieved from https://magesblog.com/post/2012-06-12-transforming-subsets-of-data-in-r-with/
@misc{ 2012-transforming-subsets-of-data-in-r-with-by-ddply-and-data.table,
author = { Markus Gesmann },
title = { Transforming subsets of data in R with by, ddply and data.table },
url = { https://magesblog.com/post/2012-06-12-transforming-subsets-of-data-in-r-with/ },
year = { 2012 }
updated = { Jun 12, 2012 }
}